Showing posts with label Protein thermal stability. Show all posts
Showing posts with label Protein thermal stability. Show all posts
Friday, February 22, 2019
The fifth sense of a protein: Quinary interactions in SOD1
The interior of living cells is an eXtremely crowded environment, with a large part of the volume being occupied by diverse macromolecules. For a protein it is like to move in a suburban train at rush hour! However, how this crowding affects the life of a protein, in particular its stability, is still unknown. The group of our collaborator (S. Ebbinghaus at the TU in Braunschweig, Germany) using rapid laser-induced temperature jumps, showed that weak transient interactions with the surrounding macromolecules, often referred to as quinary interactions (aka the fifth order of structural organization of a protein), can indeed amplify and even reverse the stability response of proteins to single-point mutations. By performing innovative multi-scale molecular simulations we shed light on the microscopic origin of those interactions, providing a possible explanation of the experimentally observed stability effects. Together, the results highlight the importance of considering weak transient interactions with the intracellular environment when investigating the relationship between stability and function in vivo as well as possible pathogenic misfolding and aggregation paths. The work is published in JACS, see here.
Thursday, December 13, 2018
The force, the heat and the shear. Three weaknesses for three perturbations!
Shear unfolding chapter two. Same question: Can proteins unfold in shearing fluid flows, and to what extent? How do the tensile forces exerted by the solvent affect the protein compared to other types of external perturbations such as thermal denaturation, or directional pulling forces used in optic/magnetic tweezers or atomic force microscopy (AFM) experiments? Before the answer: conventional all-atom molecular dynamics simulations often require too much computational effort, hence, we have developed an original methodology using Lattice Boltzmann Molecular Dynamics (LBMD) and the Optimized Potential for Efficient peptide folding Prediction (OPEP) coarse-grained model to inquire the unfolding features of a small Cold Shock Protein subjected to three different perturbations: shear flow, heat shock and pulling force. Since the implicit-solvent OPEP model inherently lacks hydrodynamics, the Lattice Boltzmann framework allowed us to realistically simulate the flow interaction with the protein, while retaining good computational efficiency with respect to explicit-solvent approaches. Here the answer: The direct comparison of the unfolding mechanisms evidenced that the three perturbations act on different weaknesses of the protein, and thus lead on average to very different unfolding pathways. Funny enough, for this small globular protein shear flow acts more similarly to thermal excitation then a direct mechanical force. Our results suggest that the interpretation of experimental studies that rely on force-spectroscopy techniques to investigate natural shear-activated systems, such as the von Willebrand factor or the bacterial adhesin FimH, is not straightforward. The paper is out here.
Wednesday, November 22, 2017
Mechanics of thermal adaptation
Somero’s corresponding state principle relates protein enhanced thermal stability with mechanical rigidity. A natural way to test the mechanical stiffness of a protein is to apply a directional force as in single molecule AFM experiments. Recent experiments have been therefore inspired by the possible correlation among the mechanical and thermal stabilities. Unfortunately single molecule experiments lack molecular resolution, and in silico realisation of thermal and mechanical unfolding can provide very useful insights. This is exactly what we have done in a recent work focused on two homologues belonging to the Cold Shock Protein family. Our results show that for these species there is not a correlation among the thermal resistance of the thermophilic Csp and its mechanical stability. The paper is out in JPC Letter [here].
Monday, September 4, 2017
Protein melting: the Lindemann criterion applied to proteins
The full knowledge of the structural and dynamic properties of the native, transition, intermediate, and denatured states of proteins is a key goal of life and physical sciences. Indeed, this kind of information can lay the foundation for the deep understanding of folding and unfolding of proteins which, in turn, are crucial processes in the metabolism of cells, regulating biological activity and targeting proteins to different cellular locations. In our work (Katava et. al., PNAS (2017) here)we focus the attention on the sub-nanosecond dynamics of a model protein in correspondence of the melting transition, when it is embedded in three different solvents. We show that, although the different solvents modify the protein melting temperature, a common scaling toward a constant value for the local fluctuations is attained when approaching the unfolding temperature.
Quite remarkably, this result is reminiscent of the famous criterion for melting of solids proposed by F.A. Lindemann in 1910, which states that crystals liquefy when their atomic root mean square fluctuations exceed a certain threshold value. The common scaling we found for protein mean square displacements at melting not only sheds unique light on the relationship between protein flexibility and stability, but also opens the possibility to predict protein unfolding in special environments (e.g., the cell interior) by following thermal local fluctuations.
The striking analogy between the melting of inorganic crystals and native biomolecules suggests that these seemingly very different systems may share similar behavior in correspondence of phase transitions. On these grounds, we may speculate that simplified theories of solids can also be effectively applied to interpret the behavior of complex biological systems.
Quite remarkably, this result is reminiscent of the famous criterion for melting of solids proposed by F.A. Lindemann in 1910, which states that crystals liquefy when their atomic root mean square fluctuations exceed a certain threshold value. The common scaling we found for protein mean square displacements at melting not only sheds unique light on the relationship between protein flexibility and stability, but also opens the possibility to predict protein unfolding in special environments (e.g., the cell interior) by following thermal local fluctuations.
The striking analogy between the melting of inorganic crystals and native biomolecules suggests that these seemingly very different systems may share similar behavior in correspondence of phase transitions. On these grounds, we may speculate that simplified theories of solids can also be effectively applied to interpret the behavior of complex biological systems.
Monday, January 30, 2017
Evolution and Thermoadaptation in Enzymes
 How
have enzymes evolved since life appeared on Earth? What has driven
the adaptation of enzyme catalysis to different temperatures?
Although massive work has been done, in 2017 these are still “hot”
questions searching for answers. An interesting paper by Nguyen et
al., just appeared on Science,
tackled this unsolved issue by investigating the molecular mechanisms
underlying thermoadaptation of enzyme catalysis through ancestral
sequence reconstruction spanning 3 billion years of evolution, and
using as a study-case the adenylate kinase (Adk). The authors assumed
as true the well-supported hot-start hypothesis, which implies that
life adapted to cooler temperatures because of the Earth’s cooling.
According to this, a thermophilic enzyme had to adapt to maintain a
high catalytic activity even at lower temperatures, while
accommodating relaxed selection on thermostability. It has been
hypothesized that enzymes overcame this thermal kinetic hurdle by
reducing the enthalpic activation barrier. However, Nguyen et
al., by reconstructing eight nodes of
the Adk lineage and expressing them together with four modern Adk
enzymes, found out something different. Indeed, from the analysis of
the Eyring plots, they showed that the oldest ancestors had a
strongly negative change in heat capacity of activation,
which can explain their extreme slow catalysis at low temperatures.
Conversely, along the thermoadaptation process toward cooler
temperatures, this kinetic obstacle has been progressively removed,
bringing the heat capacity of activation to zero. This close to zero heat capacity of activation was also observed for thermophilic enzymes evolved from mesophilic ancestors, but not for modern hyperthermophiles that remained thermophilic throughout their evolutionary pathway. This represents also a prove of the "evolutionary memory" of enzymes. To find out more about this new scenario, see here for the full manuscript.
How
have enzymes evolved since life appeared on Earth? What has driven
the adaptation of enzyme catalysis to different temperatures?
Although massive work has been done, in 2017 these are still “hot”
questions searching for answers. An interesting paper by Nguyen et
al., just appeared on Science,
tackled this unsolved issue by investigating the molecular mechanisms
underlying thermoadaptation of enzyme catalysis through ancestral
sequence reconstruction spanning 3 billion years of evolution, and
using as a study-case the adenylate kinase (Adk). The authors assumed
as true the well-supported hot-start hypothesis, which implies that
life adapted to cooler temperatures because of the Earth’s cooling.
According to this, a thermophilic enzyme had to adapt to maintain a
high catalytic activity even at lower temperatures, while
accommodating relaxed selection on thermostability. It has been
hypothesized that enzymes overcame this thermal kinetic hurdle by
reducing the enthalpic activation barrier. However, Nguyen et
al., by reconstructing eight nodes of
the Adk lineage and expressing them together with four modern Adk
enzymes, found out something different. Indeed, from the analysis of
the Eyring plots, they showed that the oldest ancestors had a
strongly negative change in heat capacity of activation,
which can explain their extreme slow catalysis at low temperatures.
Conversely, along the thermoadaptation process toward cooler
temperatures, this kinetic obstacle has been progressively removed,
bringing the heat capacity of activation to zero. This close to zero heat capacity of activation was also observed for thermophilic enzymes evolved from mesophilic ancestors, but not for modern hyperthermophiles that remained thermophilic throughout their evolutionary pathway. This represents also a prove of the "evolutionary memory" of enzymes. To find out more about this new scenario, see here for the full manuscript. Tuesday, March 8, 2016
Corresponding state hypothesis along the GTPase cycle in homologues
Marina Katava et al. have tracked the effect of substrate binding on the conformational flexibilities of two homologous GTPase domains of different stability contents by mimicking the catalytic cycle. The notable finding is that for the hyperthermophilic specie only at its high working temperature the release of entropy in the domain upon the hydrolysis of the GTP molecule matches that of the mesophilic domain at ambient condition. This was probed following several functional modes of the protein considered important for signalling propagation upon reaction as well as for the allosteric activation. It was also confirmed that the key region ensuring the flexibility for the conformational change upon catalysis (the switch I region) is also the weakest part in the mesophilic domain, confirming a sort of stability/function trade-off. You can enjoy the paper here.
Monday, February 1, 2016
Kinetic extreme stability
I was recently attracted by a paper in PNAS, "Designed protein reveals structural determinants of extreme kinetics stability", see paper here. Generally, when thinking to protein stability my first vision relates to the stability curve measuring the free energy difference between the folded and unfolded states as it changes with temperature. This is stability in thermodynamic sense. And this means, for example, in a probabilistic perspective, how many folded proteins are found at a given thermodynamic condition in a solution, and assuming that all the possible states (folded and unfolded) are sampled. But, one protein can survive in its folded state just because the barrier separating it to the unfolded floppy configuration is too high. This is the case for example for the protein a-lytic protease, that is really kinetically stable although the unfolded state is lower in energy. The authors, by de nove design constructed a protein which is at once thermodynamically and kinetically stable. The protein is named ThreeFoil and is a repeat of three peptide domains with both alpha and beta secondary structures. The interesting part of the paper is the correlation found among the extreme kinetic stability and some metrics describing the relative contacts between the amino-acids in the structure (absolute contact order, ACO, and long-range order, LRO) which in turn account for the possibility to have unfolding event highly cooperative.
| From the article in PNAS (PNAS 2015 112 (47) 14605-14610) |
Tuesday, November 17, 2015
Over 100°C
 What factors contribute to protein stability at very extreme temperatures? What gain comes from entropy and enthalpy? And how to account for the delicate effect of temperature on molecular interactions like the hydrophobic and ionic ones? All this is tackled in a very intriguing work by Y. Matsura et al. "Thermodynamics of protein denaturation at temperature over 100°C: CutA1 mutant proteins substituted with hydrophobic and charged residues" recently published in Scientific Reports [see here]. By designing sequential mutations the authors were able to construct hyper-stable versions of the CutA1 protein and to extract the main thermodynamic parameters characterising their thermal stability. It is a very important work challenging both technical biochemical problems, like the aggregation of proteins generally occurring above 80°C, and the basic thermodynamics controlling protein stability over 100°C where for instance hydrophobic interactions cease to be entropically driven and ion-pairing can benefit from water dielectric constant decrease.
What factors contribute to protein stability at very extreme temperatures? What gain comes from entropy and enthalpy? And how to account for the delicate effect of temperature on molecular interactions like the hydrophobic and ionic ones? All this is tackled in a very intriguing work by Y. Matsura et al. "Thermodynamics of protein denaturation at temperature over 100°C: CutA1 mutant proteins substituted with hydrophobic and charged residues" recently published in Scientific Reports [see here]. By designing sequential mutations the authors were able to construct hyper-stable versions of the CutA1 protein and to extract the main thermodynamic parameters characterising their thermal stability. It is a very important work challenging both technical biochemical problems, like the aggregation of proteins generally occurring above 80°C, and the basic thermodynamics controlling protein stability over 100°C where for instance hydrophobic interactions cease to be entropically driven and ion-pairing can benefit from water dielectric constant decrease. Sunday, November 1, 2015
Water helps life in extreme environments?
A few months ago we published an explorative work focusing on the possible contribution of water molecules buried in the interior of proteins to their different thermal stabilities [see here for the paper]. The study-case was a pair of homologous GTPase domains from a mesophilic and a hyperthermophilic organism, respectively. Now, we extended our approach by considering a large set of homologous pairs. Let's list the main findings. Firstly, for some homologues internal water gives a meaningful contribution to the stability gap in favour of the thermophilic variant. This was probed at ambient condition. Secondly, when considering the behaviour at high temperature, we found that thermophilic proteins are more keen to maintain their internal cavities wet, and therefore benefiting by this wetting. We propose that internal hydration can be viewed as an alternative tuneable variable for the engineering of proteins with enhanced stability. Enjoy the manuscript here.
Thursday, October 8, 2015
Fast and Cheap: the bioinformatics help to membrane protein thermal-stabilisation!
A very interesting work by Sauer et al. has been recently published in Biophysical J (here) and was highlighted by a nice comment by CG Tate (see here). What the story is about? Sauer and colleagues have deployed comparative bioinformatics tools to propose single point mutations for stabilising a membrane protein, the tetracycline antiporter from Bacillus subtilis (BsTetL). Out of the 22 mutations tested, 7 were proved to increase the thermal stability of the protein: almost 32% of success! This is a very high number. Why all this interest? Following the comment by Tate, membrane proteins -that are key targets for many drugs- are difficult to be crystallised. They must be isolated from the membrane environment and solubilised before crystallised. This is achieved by using detergents that protect the hydrophobic surface of the membrane protein from the contact with water. The stability of the protein/detergent complex is considered a pre-requisite for the success of the crystallisation process. While many effort has been placed to select/design the correct detergent to envelope the membrane protein, and alternative strategy consists in making the protein more stable. But select what to mutate, because of the highly complicate procedure, is essential. In this regards, the methodology proposed by Sauer et al. could be a breakthrough. Of course, questions remain. From the long research on thermophilic proteins we know that often stability trades functionality. Ergo, making the protein more stable could lock it in not functional states. This concerns is well explained in the Tate's comment. Go there, for an elegant discussion.
Thursday, September 24, 2015
In vivo stability! Cell type matters.....
In a very recent PNAS, J. Danielsson et al. [see here] present a very interesting work focusing on protein stability in cell. They used in-cell NMR to reconstruct the stability curve of the protein SOD1. The interesting finding is that when the protein is moved in two different types of cells, a bacterial (E. coli) and a mammalian cell (A2780) the protein is destabilised in both cases. Firstly, this finding questions the common believe that under crowding a protein gets stabilised because of an excluded volume effect. In short, if the available space is reduced because of the presence of a large numbers of macromolecules acting as crowders, the highly entropic and extended unfolded state should be unfavored. This picture is however very simplified since in both folded and unfolded states, a targeted protein interacts with its neighbours, and the results of these specific interactions, i.e. electrostatic, could alter the equilibrium favouring unfolding. The effect of different specific interactions, is actually probed by the authors, showing that by changing the local environment, moving from E.coli to a mammalian cell, the destabilisation effect is different. Last, but not least, the destabilisation results as an increase of the specific heat of unfolding that shrinks the stability curve. This calls for a particular effect of the crowders on the nature of the unfolded state. Stay tune, because the life of proteins in cell is where our interest is going....
| Molecular view of molecular crowding. Project at the Riken HPC center, Japan [see here] |
Monday, May 18, 2015
Designing thermal stability via non-equilibrium simulations
The design of protein thermal stability is appealing for practical uses.
In a recent work [1] by Tian, Woodard, Whitney and Shakhnovich [see here] non-equilibrium Monte Carlo simulations were
effectively used to explore mutations of the Dihydrofolate Reductase
(DHFR) and their impact on both the stability and functionality of the
enzyme.
The key point of the work is the use of non-equilibrium Monte Carlo (MC) simulations. A protein is excited at different temperatures and its
"unfolding " as function of MC steps is recorded.
For each temperature, the average value of a given observable or order
parameter that describes the state of the protein, ie the root mean
square displacement with respect to the native state, the energy, the
gyration radius, depends on the simulation length -in the specific case
the number of MC steps. This relates to the fact that the transition
from the folded to the unfolded state, for a given temperature, is rate
limited by the free energy barrier dividing the two states. How this
dependence can be washed up when considering the effect of mutations?
The authors proved a nice recipe: first, a mutation affects the
thermodynamics of the system, formally the free energy difference
between folded and unfolded state, but also the kinetics for the
folded/unfolded transition, aka the free energy barrier dividing the two
state. It is possible to image that the thermodynamic effect is
mirrored on the change of free energy barrier via a scaling factor that measures
how the mutation influences the transition state of the folding/unfolding process.
Secondly, when considering the non-equilibrium MC simulations for both the
wild type and the mutant, the shift of the apparent melting temperature
(the temperature leading unfolding) of the mutated system with respect
to WT results independent from the simulation length. This can be formally showed, and the reader is invited to dig the work.
Using this strategy several mutations stabilizing the protein and that
maintain functionality were identified. I wonder whether this approach
can be used straightforwardly also for estimating the effect of
mutations on mechanical stability.
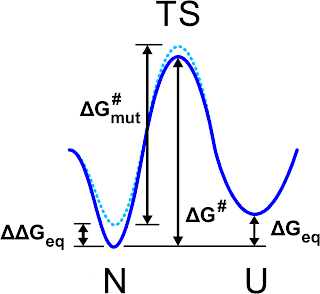 |
| Schematic view of the free energy profile for the folded (N) unfolded (U) states as it is pictured in Fig. 1 of Ref. 1 |
[1] J. Tian, JC Woodard, A. Whitney, EI Shakhnovich, Plos Comp Bio (2015) 11, e1004207.
Wednesday, December 3, 2014
Interface matters: The stiffness route to stability of a thermophilic tetrameric malate dehydrogenase
Understanding the relation between protein flexibility, stability and function remains one of the most challenging, open questions in biophysical chemistry. For example, proteins need to be flexible to facilitate substrate binding but locally rigid to sustain substrate specificity. Enzymes from microorganisms that thrive at elevated temperatures, also referred to as thermophiles, are a natural study-case to dig into the issue. These proteins are stable and functional at a high temperature regime but generally lack activity at ambient conditions. Therefore, their thermal stability has been correlated to enhanced mechanical rigidity through the so-called corresponding states paradigm introduced years ago by Somero. The generality of this view, however, has been questioned by a number of experimental and computational studies. Computer simulations based on the molecular dynamics technique offer a unique opportunity to explore the correlation among mechanical rigidity and thermophilicity. In our recently published article in PLoS One we consider the specific case of two tetrameric orthologous malate dehydrogenase proteins from two bacteria that grow optimally at different temperatures. For these orthologues, as for other oligomeric proteins, the role of interfacial interactions becomes critical, adding up to the other cohesive forces acting on monomeric proteins. How the protein rigidity/flexibility patterns influence the stability and function of the two molecules is discussed in detail in the paper.
Subscribe to:
Posts (Atom)






